2025 Student Poster Session
On Friday, November 14, 2025, the department hosted its annual Student Poster Session, featuring 15 students presenting their ongoing research which was hosted in the SAS Hall Commons. The event brought together faculty, graduate students, and undergraduates who came to browse an array of projects while enjoying bagels, coffee, and conversations. The lively atmosphere showcased not only the students’ hard work but also the collaborative spirit that defines our program.
Below, we highlight each participating student along with their title and abstract. See the full event gallery here.

Jingtian Bai
Title: Feature-Based Transfer Learning with Integrated False Negative Control
Abstract: Transfer learning has become a pivotal approach in machine learning, facilitating the transfer of knowledge from one domain to enhance predictive accuracy in another. This capability is particularly vital in applications where the target domain often suffers from limited labeled data or challenging feature spaces. Despite its promise, the practical deployment of transfer learning faces a critical challenge: heterogeneity across domains. Failing to address variations in data distributions, feature representations, and noise levels can lead to significant biases, resulting in suboptimal model performance. In this work, we present a novel transfer learning framework tailored for high-dimensional regression analysis. The framework incorporates high-dimensional false negative control (HD-FNC) to effectively retain signal variables while mitigating the influence of noise and domain-change-induced biases. Importantly, HD-FNC performs efficient dimension reduction by distinguishing noise variables from weak true signals. By combining HD-FNC with retraining strategies that adapt Ridge and Ridgeless regression models to target populations, our method ensures robust signal detection, improved predictions, and enhanced generalization.

Timothy Boakye
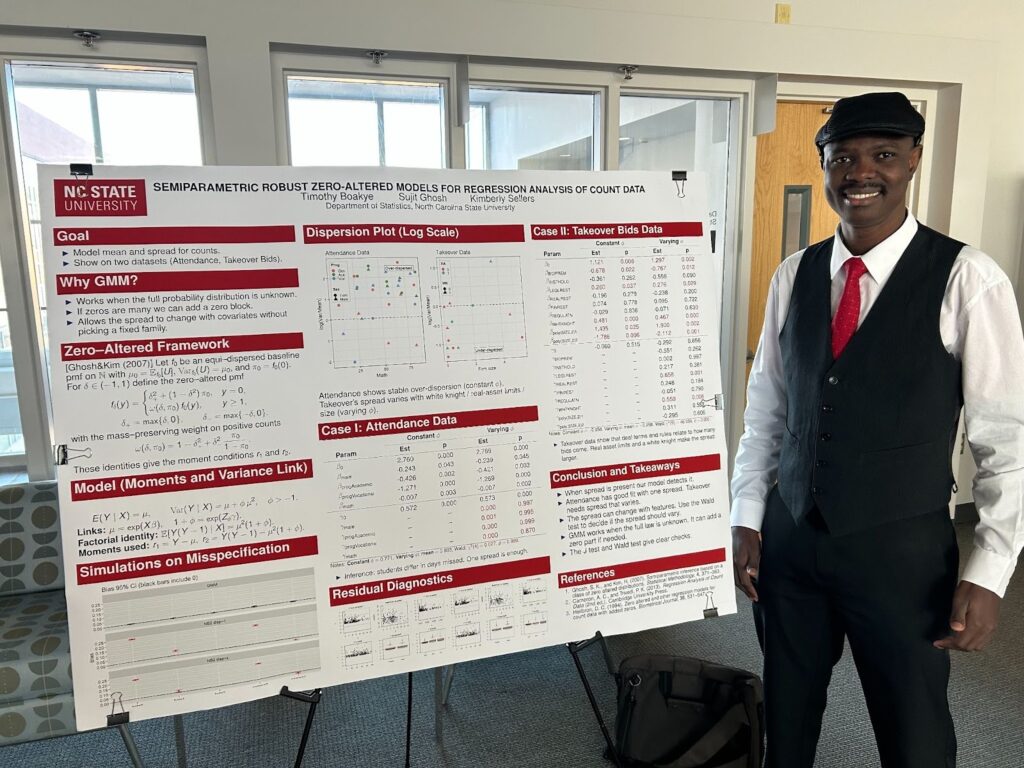
Title: Semiparametric Robust Zero-Altered Models for Regression Analysis of Count Data
Abstract: We propose a flexible class of semiparametric regression models for count data based on zero-altered distributions, estimated via the generalized method of moments (GMM). Unlike conventional parametric generalized linear models (GLMs; e.g., Poisson or negative binomial), our approach accommodates both under- and over-dispersion while retaining full support on the nonnegative integers. By relying only on moment conditions rather than full likelihoods, the GMM framework offers robust inference even under distributional misspecification, and ensures consistent estimation when higher-order features are misspecified. Theoretical results establish consistency and asymptotic normality of the estimators under broad conditions. The robustness and flexibility of the proposed models are extensively demonstrated using three real-world data sets covering diverse dispersion patterns and practical domains, which highlight superior performance relative to standard GLMs. Accompanying R functions facilitate straightforward implementation for applied researchers.

Rishav Chakrabarti
Title: Latent Network Bayesian Regression for Functional Neuroconnectivity
Abstract: Understanding how brain regions interact at rest is key to uncovering neural mechanisms underlying conditions such as autism. However, analyzing resting-state fMRI data is computationally challenging due to the massive number of voxel-level connections. In this project, we propose Latent Network Bayesian Regression (LNBR), a Bayesian regression model for network response data that introduces a second, latent network as a covariate, while simultaneously including individual characteristics such as age, IQ, and autism status. For computational efficiency, we approximate the shortest-path distance matrix implied by our latent network using a small set of landmark nodes (voxels). We complete our hierarchical model with priors that promote both sparsity and interpretability among the individual latent networks. We use an adaptive Gibbs sampler to ensure efficient inference. Simulation studies show that LNBR accurately recovers underlying connectivity patterns, identifies informative landmarks, and
reveals meaningful associations between brain network organization and behavioral traits. This framework provides a scalable and interpretable approach for studying individualized brain connectivity in large-scale fMRI datasets.

Chris Chung
Title: Change Point Detection in Correlated Time-Varying Networks
Abstract: In the context of offline change point detection, we study a sequence of inhomogeneous Erdős–Rényi graphs with arbitrary dependence structures. From the assumption that the edges follow a marginally Bernoulli distribution, the probability generating matrices are constant over time except at unknown change points. This setup extends the dependence from the latent positions in existing literature without any assumptions on the dependence of edges.
To detect these change points, we propose a sequential Frobenius norm statistic for the general case and a GLS weighted statistic for the AR(1) correlation structure. In both statistics, the Frobenius norm difference of adjacency spectral embeddings are thresholded by a detection lower bound that is minimax optimal for both dependent and independent cases. While the sequential statistic compares consecutive adjacency matrices, the GLS method computes a weighted average of adjacency matrices from the previous change point. Consistent change point estimation has also been proven theoretically with numerical examples in both simulated and real data settings showing competitive performance. Extensions to network structures of arbitrary rank are also discussed.
Adira Cohen
Title: Don’t Throw Away Uncertainty: Using All AI Confidence Scores to Model Deer Body Condition
Abstract: Camera traps have proven themselves to be an efficient and effective approach for studying a wide range of ecological processes. Camera traps are remote collection devices which capture images of animals as they move across the field of view, allowing the collection of large image datasets with relatively little human effort. The size of the datasets creates a bottleneck when translating the images into usable information, which has recently been tackled by integrating AI into the workflow. However, the data procured from AI approaches are of a different nature than that obtained from humans processing the images (i.e., human annotations), necessitating new statistical methods in order to obtain inference, make predictions, and quantify uncertainty. We propose a Bayesian hierarchical data-fusion model which combines the strengths of human annotations and AI predictions. The benefits of our approach are an ability to provide uncertainty quantification as well as improved inference and prediction power. Our analysis uses camera trap data from the Candid Critters project, which encompasses all of North Carolina. We are interested in using the images from this study to understand the relationship between a metric for health (namely, body condition) of white-tailed deer and their environment. We conduct a simulation study which demonstrates the improved inference and uncertainty quantification of our method and apply our model on the Candid Critters dataset to give novel ecological inference.
Prithwish Ghosh
Title: Automated Sliced Gibbs sampling for Arbitrary Kernels
Abstract: We introduce the Automated Sliced Gibbs (ASG) sampler — an adaptive and fully automated MCMC framework for sampling from arbitrary multivariate probability kernels. ASG combines a Cauchy-based effective support estimation with slice-driven Gibbs updates, removing the need for manual tuning of bounds or proposals. The method preserves the invariance and ergodicity of classical Gibbs sampling while efficiently handling complex, multimodal, and non-Gaussian targets. Theoretical results establish stationarity, and simulations on Gaussian mixtures, Rosenbrock, Ackley, LASSO, and Hinge-loss kernels show that ASG attains higher effective sample size per second and faster mixing than Random-Walk Metropolis–Hastings, providing a robust, scalable solution for high-dimensional sampling.

Carter Hall
Title: Quantifying Uncertainty in a Mathematical Model of Autonomic Dysfunction
Abstract: Autonomic dysfunction involves impaired control of involuntary physiological processes such as heart rate, blood pressure, and respiration, contributing to conditions like diabetes, Parkinson’s disease, and chronic pain. Yet, no automated or quantitative diagnostic tools currently exist. We propose a novel mathematical model of the Valsalva Maneuver—a noninvasive test that elicits autonomic responses—integrating blood pressure and ECG-derived measures of heart rate and respiration. The model parameters capture key physiological mechanisms. Instead of traditional Fisher information–based sensitivity analysis, which can be unreliable for nonlinear or correlated parameters, we use Hamiltonian Monte Carlo to obtain the full joint posterior distribution of parameters. This approach is extended across multiple patients to yield individualized posterior estimates, which are further analyzed through regression on covariates such as age and gender. This two-stage hierarchical analysis aims to explain additional variability and guide refinement of the underlying physiological model.

Sohyeon Kim
Title: Multivariate Multinomial Logit Model with ANOVA Decomposition for Correlated Categorical Outcomes
Abstract: In many medical studies, it is often necessary to collect multiple categorical outcomes that are interdependent. However, most existing research on multinomial regression fits each outcome separately, which ignores correlations between outcomes, and thus may lead to loss of information and reduced predictive accuracy. Accounting for correlations between multilevel categorical outcomes requires a high-dimensional parameter space, making model estimation challenging. In this paper, we propose a multivariate multinomial logit model that captures outcome correlations and reduces the parameter space using ANOVA decomposition. The ANOVA decomposition enables explicit conditional model formulations, which allow for a computationally much simpler composite likelihood approach. We develop an efficient Minorization-Maximization algorithm to maximize the composite likelihood, which also incorporates variable selection via a bridge penalty. Simulation studies are conducted to evaluate our method, demonstrating its effectiveness in parameter estimation and variable selection. We further illustrate our method using data from the AURORA study.

Arpan Kumar
Title: Predictive Subsampling for Scalable Inference in Networks
Abstract: Network datasets appear across a wide range of scientific fields, including biology, physics, and the social sciences. To enable data-driven discoveries from these networks, statistical inference techniques like estimation and hypothesis testing are crucial. However, the size of modern networks often exceeds the storage and computational capacities of existing methods, making timely, statistically rigorous inference difficult. In this work, we introduce a subsampling-based approach aimed at reducing the computational burden associated with estimation and two-sample hypothesis testing. Our strategy involves selecting a small random subset of nodes from the network, conducting inference on the resulting subgraph, and then using interpolation based on the observed connections between the subsample and the rest of the nodes to estimate the entire graph. We develop the methodology under the generalized random dot product graph framework, which affords broad applicability and permits rigorous analysis. Within this setting, we establish consistency guarantees and corroborate the practical effectiveness of the approach through comprehensive simulation studies.
Ruohan Liao
Title: A Causal Mediation Framework for Multiple Exposures and Multiple Mediators
Abstract: This research develops a causal mediation analysis framework that integrates Directed Acyclic Graph (DAG) learning with Linear Structural Equation Modeling (LSEM) to quantify individual-level causal effects across multiple exposures and correlated mediators. Motivated by understanding how maternal environmental exposures jointly influence newborn outcomes through epigenetic mechanisms, this framework generalizes traditional mediation decomposition to complex multi-exposure, multi-mediator networks and enables interpretable path-specific effect estimation.
Building upon the structural relationships encoded by the DAG, we derived a general class of estimators that capture hierarchical causal pathways among exposures, mediators, and outcomes, linking each exposure to the outcome through both intermediate exposures and mediators. This formulation allows for interpretable effect decomposition at the individual level, accommodates correlated mediators, and captures inter-exposure dependencies. Furthermore, we establish the asymptotic properties of these estimators to enable valid inference. Applied to data from the Newborn Epigenetics Study (NEST) cohort, this framework provides new insights into how maternal exposures influence birth outcomes through coordinated epigenetic alterations. Overall, our work extends traditional mediation analysis to a multivariate, graph-based context, offering a flexible and theoretically grounded approach for uncovering complex biological pathways.

Yi Liu
Title: COADVISE: Covariate Adjustment with Variable Selection in Randomized Controlled Trials
Abstract: Adjusting for covariates in randomized controlled trials can enhance the credibility and efficiency of treatment effect estimation. However, handling numerous covariates and their complex (non-linear) transformations poses a challenge. Motivated by the case study of the Best Apnea Interventions for Research (BestAIR) trial data from the National Sleep Research Resource (NSRR), where the number of covariates (p=114) is comparable to the sample size (N=196), we propose a principled Covariate Adjustment with Variable Selection (COADVISE) framework. COADVISE enables variable selection for covariates most relevant to the outcome while accommodating both linear and nonlinear adjustments. This framework ensures consistent estimates with improved efficiency over unadjusted estimators and provides robust variance estimation, even under outcome model misspecification. We demonstrate efficiency gains through theoretical analysis, extensive simulations, and a re-analysis of the BestAIR trial data to compare alternative variable selection strategies, offering cautionary recommendations. A user-friendly R package, Coadvise, is available to facilitate practical implementation.

Ayumi Mutoh
Title: Pitfalls and remedies for penalized maximum likelihood estimation of Gaussian processes
Abstract: Gaussian processes (GPs) are popular as nonlinear regression models for expensive computer simulations, yet GP performance relies heavily on estimation of unknown covariance parameters. Maximum likelihood estimation (MLE) is common, but it can be plagued by numerical issues in small data settings. The addition of a nugget helps but is not a cure-all. Penalized likelihood methods may improve upon traditional MLE, but their success depends on tuning parameter selection. We introduce a new cross-validation (CV) metric called ”decorrelated prediction error” (DPE), within the penalized likelihood framework for GPs. Inspired by the Mahalanobis distance, DPE provides more consistent and reliable tuning parameter selection than traditional metrics like prediction error, particularly for K-fold CV. Our proposed metric performs comparably to standard MLE when penalization is unnecessary and outperforms traditional tuning parameter selection metrics in scenarios where regularization is beneficial, especially under the one-standard error rule.

Sihyung Park
Title: Evaluating and Learning Optimal Dynamic Treatment Regimes under Truncation by Death
Abstract: Truncation by death, a prevalent challenge in critical care, renders traditional dynamic treatment regime (DTR) evaluation inapplicable due to ill-defined potential outcomes. We introduce a principal stratification-based method, focusing on the always-survivor value function. We derive a semiparametrically efficient, multiply robust estimator for multi-stage DTRs, demonstrating its robustness and efficiency. Empirical validation and an application to electronic health records showcase its utility for personalized treatment optimization.

James Robertson
Title: Prediction from possibilistic inferential models (IMs) applied to Gaussian mixtures
Abstract: Statistical inference in the style of Martin and Liu, referred to commonly as inferential models (IMs), offers proven assurances regarding validity, a desirable error control property only achievable through the use of imprecise probabilities. While this validity property is well-established in inference, there still remains work to be done in regard to valid prediction. This work explores one approach for valid prediction from a valid possibility contour, relying on a possibilistic Bernstein–von Mises theorem for its estimation via a Monte-Carlo style algorithm presented.

Ananya Roy
Title: E-values for measurement error models
Abstract: Measurement error in covariates fundamentally distorts the geometry of likelihood-based inference, leading to attenuation bias, loss of identifiability, and unstable statistical conclusions. This work investigates two foundational strategies for handling latent covariates in measurement error models – profiling and marginalization – within a unified likelihood framework. In particular, within the context of multiple testing we construct an e-value for each approach, pertaining to hypotheses about covariate effects in regression. Using the growth-rate optimality (GRO) criterion from safe testing theory, we derive analytical expressions for the expected evidence growth under each approach and establish their relative efficiency. Priors on the latent covariates emerge as a key ingredient for restoring identifiability and stabilizing inference, connecting classical regularization ideas with modern sequential optimality principles. Through both theoretical analysis and simulation, we demonstrate that profiling achieves systematically higher GRO values, indicating faster evidence accumulation. The results delineate clear regimes where one strategy dominates the other, clarifying the statistical and computational trade-offs inherent in inference under measurement error.
- Categories: